Contents
Nucleic acid structure
2'-Deoxyribonucleic acid (DNA)
DNA (2'-deoxyribonucleic acid) is the molecular store of genetic information in nearly all living systems. It is a large polymeric molecule composed of monomers known as nucleotides. Each nucleotide consists of a heterocyclic base, a pentose sugar (2'-deoxy-d-ribofuranose), and a phosphate group. There are four heterocyclic bases in DNA: adenine (A), guanine (G), cytosine (C) and thymine (T). Their structures and numbering system are shown in Figure 1.
Adenine and guanine are purines and cytosine and thymine are pyrimidines (Figure 2).
The deoxyribose sugar is shown in Figure 3. As a free sugar it can mutarotate under certain conditions, adopting furanose, acyclic and pyranose forms; but in DNA it is fixed as a furanoside.
The phosphate group can be found at the 5'- or 3'-position of the sugar depending on the method used to break down DNA to produce the nucleotides. Removal of the phosphate gives rise to a nucleoside. The heterocyclic base is linked to the 1'-position of the sugar. The chemical structures of a deoxynucleoside and a deoxynucleotide are shown in Figure 4.
The bond joining the 1'-carbon of the deoxyribose sugar to the heterocyclic base is the N-glycosidic bond. Rotation about this bond gives rise to syn and anti conformations (Figure 5). Rotation about this bond is restricted and the anti conformation is generally favoured, partly on steric grounds.
The nucleosides in the DNA duplex adopt the anti conformation (there are very few exceptions to this rule, one of which is guanosine monophosphate, in which the guanine base adopts the syn conformation about the glycosidic bond).
The 1'-position of the deoxyribose sugar is the anomeric centre. If a substituent attached to the 1'-carbon lies on the same face of the sugar ring as the 5'-hydroxyl group, it is known as the β-anomer; if the substituent is on the opposite side of the sugar ring it is the α-anomer (Figure 6). All of the nucleosides in DNA are in the β-configuration.
Ribonucleic acid (RNA)
Some organisms, for example retroviruses, use ribonucleic acid (RNA) instead of DNA as their store of genetic information. RNA is chemically very similar to DNA but there are two important differences:
- RNA has a hydroxyl group attached to the 2'-position of the sugar, and
- the pyrimidine uracil (U; Figure 7) replaces thymine in RNA.
The key biological role of RNA is as a messenger: it reads the genetic code in DNA (transcription) and transports it to the ribosome, where it is decoded into the sequence of a protein (translation).
Oligonucleotides
A dinucleotide (dimer) of DNA or RNA is formed by covalently linking the 5'-phosphate group of one nucleotide to the 3'-hydroxyl group of another to form a phosphodiester bond. An oligonucleotide (oligomer) is formed when several such bonds are made, and naturally-occurring nucleic acids are linear, high molecular weight molecules of this kind. At physiological pH (7.4) each phosphodiester group exists as an anion (hence the term nucleic acid), and nucleic acids are therefore highly charged polyanionic molecules (Figure 8).
One end of a nucleic acid strand has a 5'-hydroxyl group (primary hydroxyl) and the other end has a 3'-hydroxyl group (secondary hydroxyl). The nucleic acid chain therefore has directionality. The tetranucleotide in Figure 8a has the sequence 5'-GCAT-3' and the tetranucleotide in Figure 8b has the sequence 5'-TACG-3'. By convention, the prefixes 5'- and 3'- are not written, and nucleic acid sequences are written in the 5' to 3' direction. The oligonucleotides GCAT and TACG are distinct molecules with different chemical and biophysical properties.
Nucleic acid duplexes
The chemical structure of a single strand of DNA gives little insight into its biological function as a carrier of genetic information. However, when James Watson and Francis Crick showed in 1953 that DNA adopts a double-stranded structure (duplex), the mechanism of DNA replication (copying) became apparent. The double-helical structure was principally elucidated from X-ray fibre diffraction data (acquired by Rosalind Franklin and Maurice Wilkins) and Chargaff's rules. Erwin Chargaff discovered that the molar amount of adenine in DNA was always equal to that of thymine and the same was true for guanine and cytosine (i.e. number of moles of G = number of moles of C). Watson and Crick were able to explain this by building models to show that the two strands of DNA are held together by hydrogen bonds between individual bases on opposite strands. The purine base A always pairs with the pyrimidine T and the purine G always pairs with the pyrimidine C (Figure 9).
Not surprisingly A·T and G·C are known as Watson-Crick base pairs. They are pseudosymmetric and if an A·T base pair is laid over any other base pair (T·A, G·C or C·G) the phosphodiester backbones fall on top of each other (Figure 10 and Figure 11). Thus all four base pairs fit neatly into the double helix.
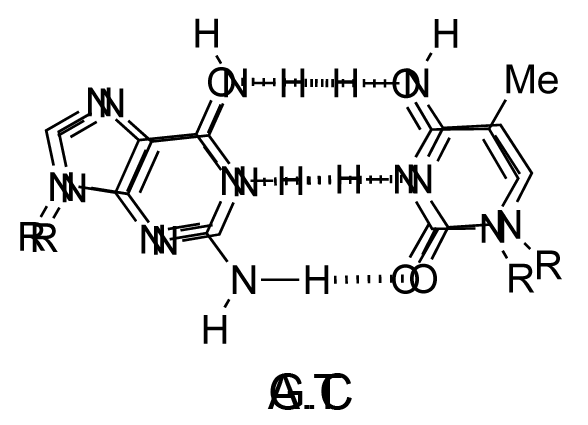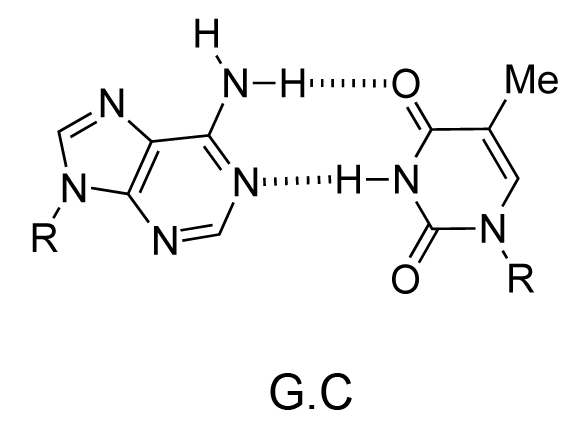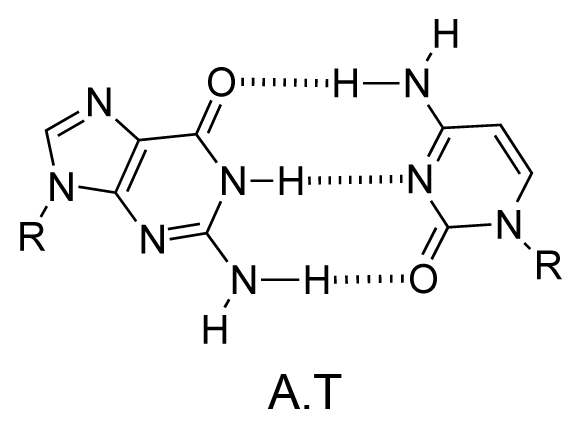
The sequence of one strand of DNA precisely defines the sequence of the other; the two strands are said to be complementary, and are sometimes called reverse complements of each other. The two strands are antiparallel, with the 5'-end of one strand adjacent to the 3'-end of the other. The two strands coil around each other to form a right-handed double helix, with the hydrophobic base pairs in the centre and the sugars and negatively charged phosphates forming the external hydrophilic backbone. The term "right-handed" indicates that the backbone at the front of the molecule facing the observer slopes down from top right to bottom left. The planar heterocyclic bases stack one on another and the separation between successive base pairs along the helix axis is around 0.34 nm. One helical turn (a full 360° turn of the double helix) is repeated every 10 to 11 base pairs. The stability of the duplex is derived from both base stacking and hydrogen bonding. The principal form of double helical DNA, B-DNA (Figure 12, middle), has a wide major groove and a narrow minor groove running around the helix along the entire length of the molecule. Proteins interact with the DNA in these grooves (principally in the major groove) and some small drug molecules (e.g. netropsin, distamycin) bind in the minor groove (see Nucleic acid-drug interactions).
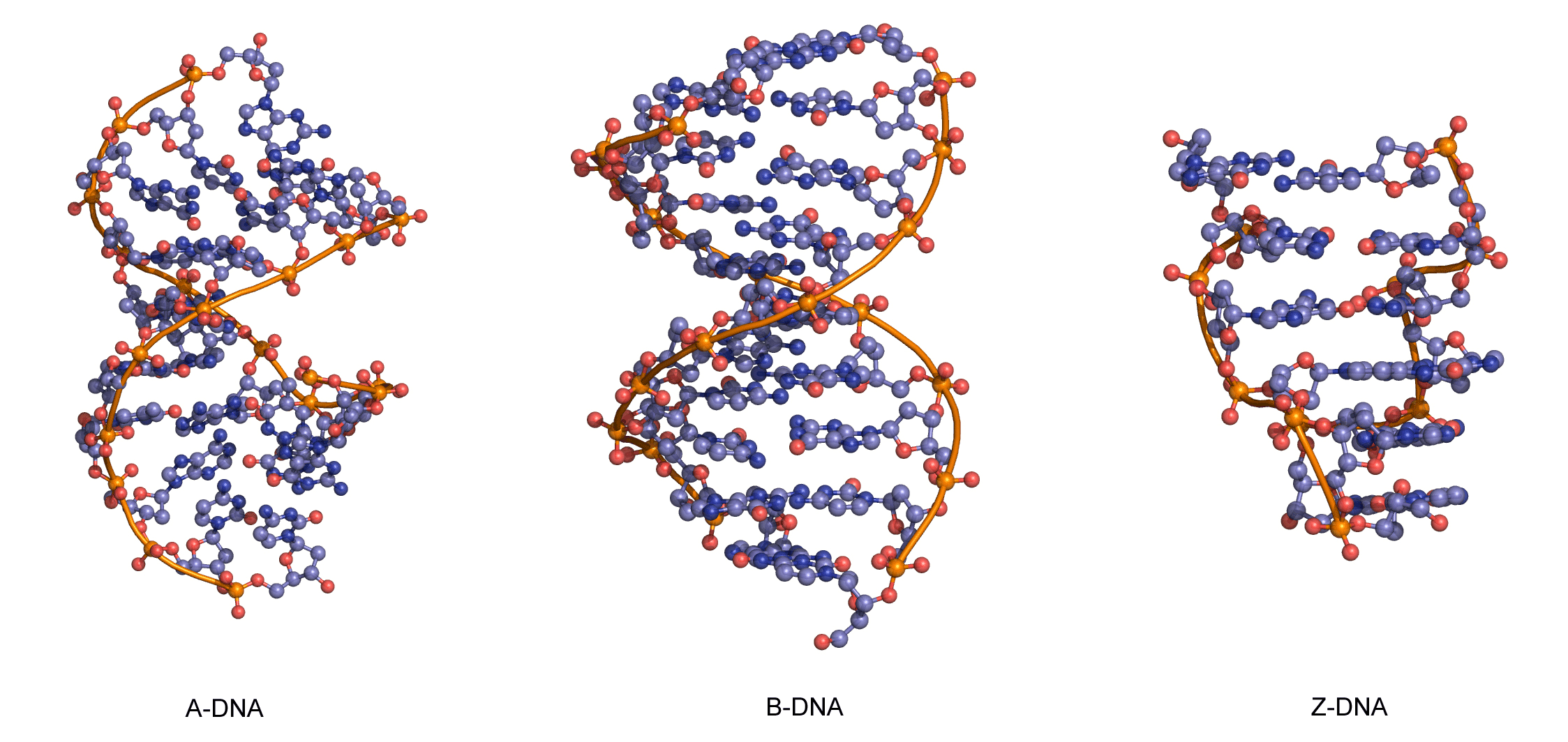
RNA can also form a right-handed duplex using the same base pairing rules (A·U and G·C), but the RNA duplex has a distinctive shape (the A-form) in which the major groove is deep and the minor groove very shallow. Under conditions of low humidity DNA can also adopt the A-form (Figure 12, left), and there are a number of other conformations of DNA and RNA, most of which are subtle variations on the A- and B-forms.
One drastically different DNA conformation, Z-DNA, (Figure 12, right) was determined from an X-ray crystal structure of a chemically synthesized DNA strand of the alternating sequence dCGCGCG, which spontaneously forms a duplex in aqueous buffer. Z-DNA is left-handed and has a dinucleotide repeat unit, so the backbone is not smooth but appears to “zig-zag”. It is not clear whether Z-DNA has biological relevance.
Techniques for determining nucleic acid structure
Nuclear magnetic resonance (NMR)
NMR is a highly developed and powerful spectroscopic technique that is valuable in the investigation of the structural, thermodynamic and kinetic properties of nucleic acids. The technique can be used to study DNA duplexes, triplexes, quadruplexes, hairpin loops, RNA duplexes and other secondary and tertiary RNA structures. Primarily, 1H NMR is used in aqueous buffers with water suppression. 31P NMR is also useful to study the environment of the nucleic acid phosphodiester backbone. NMR is most useful for studying nucleic acid structure at the local level, i.e. the nature of base pairs and triplets, nucleic acid-drug interactions and nucleic acid-protein interactions. It is less powerful at giving information on global properties (e.g. A versus B helix, DNA duplex bending) because local interactions have to be used as an indication of global properties. NMR should always be used in conjunction with other physical techniques such as circular dichroism (indicative of conformation, A, B, Z), Ultraviolet melting of duplexes, triplexes and quadruplexes (thermodynamic information), X-ray diffraction (high resolution structure analysis), and fluorescence resonance energy transfer (distance measurements). Typical NMR chemical shifts of base and sugar protons are shown in Figure 13 and Figure 14.
Crystallization and X-ray diffraction
Advances in solid-phase oligonucleotide synthesis and purification have allowed a large number of oligonucleotides to be crystallized and their molecular structures solved. This has provided important information on both the B and A forms of DNA, confirming predictions made from earlier fibre-diffraction studies, and providing high-resolution information on the sequence dependence of nucleic acid structure. X-ray diffraction of single crystals such as those shown in Figure 15 provides an accurate picture of macromolecular structure with a resolution that exceeds that of other techniques.
When a crystal is placed in the path of an X-ray source it will generate a diffraction pattern, from which, with the help of some complicated mathematics, a three-dimensional picture of the unit cell and the molecules present can be generated. X-ray crystallography requires highly ordered crystals, and obtaining suitable crystals of nucleic acids is normally the slowest part of the process. A highly concentrated solution of an oligonucleotide is required to provide a thermodynamic driving force for crystal formation. The solubility limit is exceeded by gradually removing water from the oligonucleotide solution using an external dehydrating agent, or diffusing a precipitant such as isopropanol into the solution. High-resolution structures of A, B and Z-DNA have been obtained by X-ray crystallography and the technique has been used to study DNA-drug complexes in atomic detail. In most cases the resolution of the structure does not extend beyond 2 Å (0.2 nm), but this is sufficient to provide a reasonably clear picture of the heterocyclic bases and sugar-phosphate backbone.